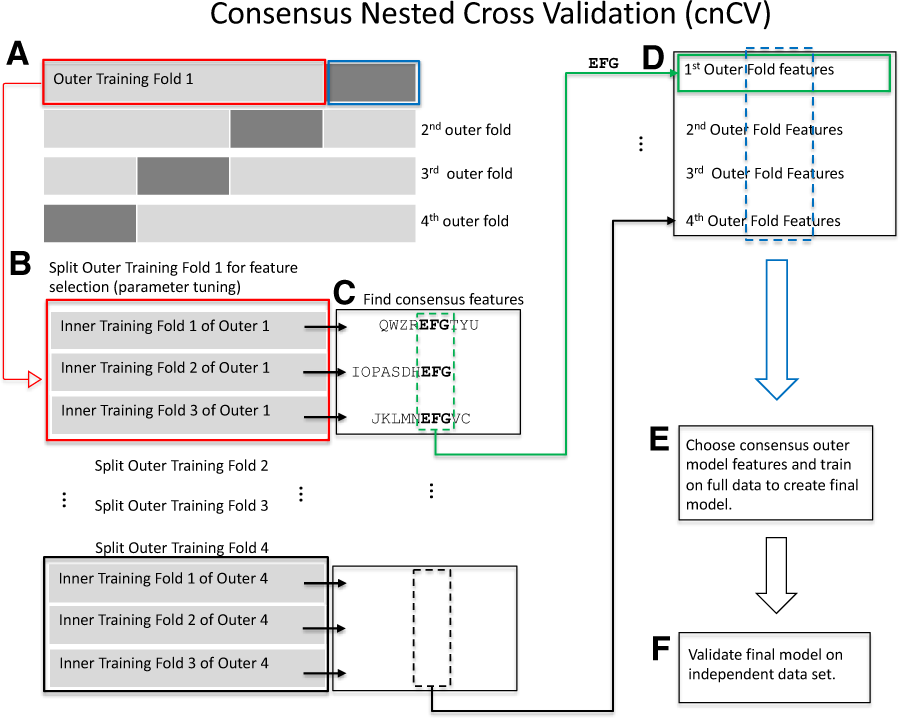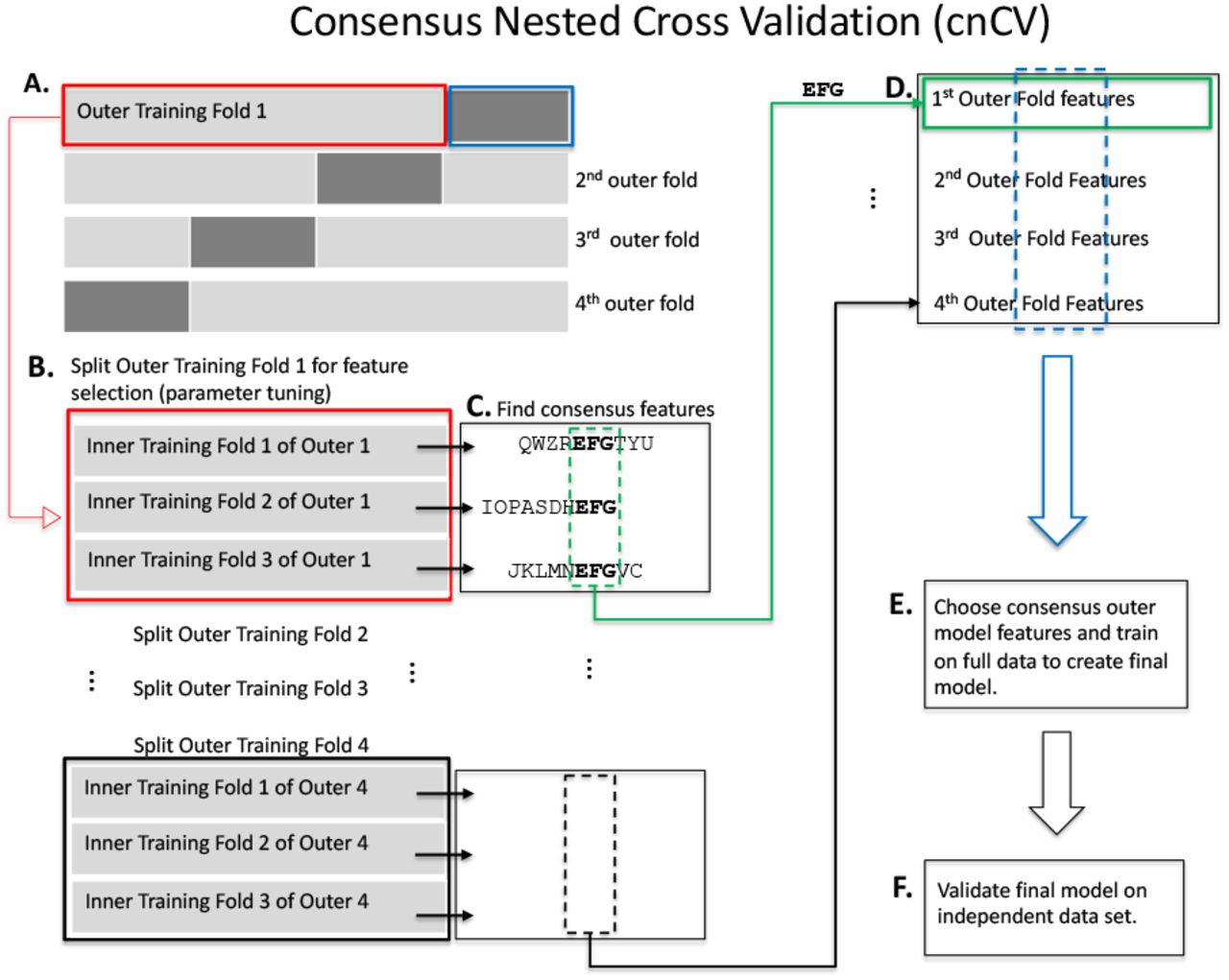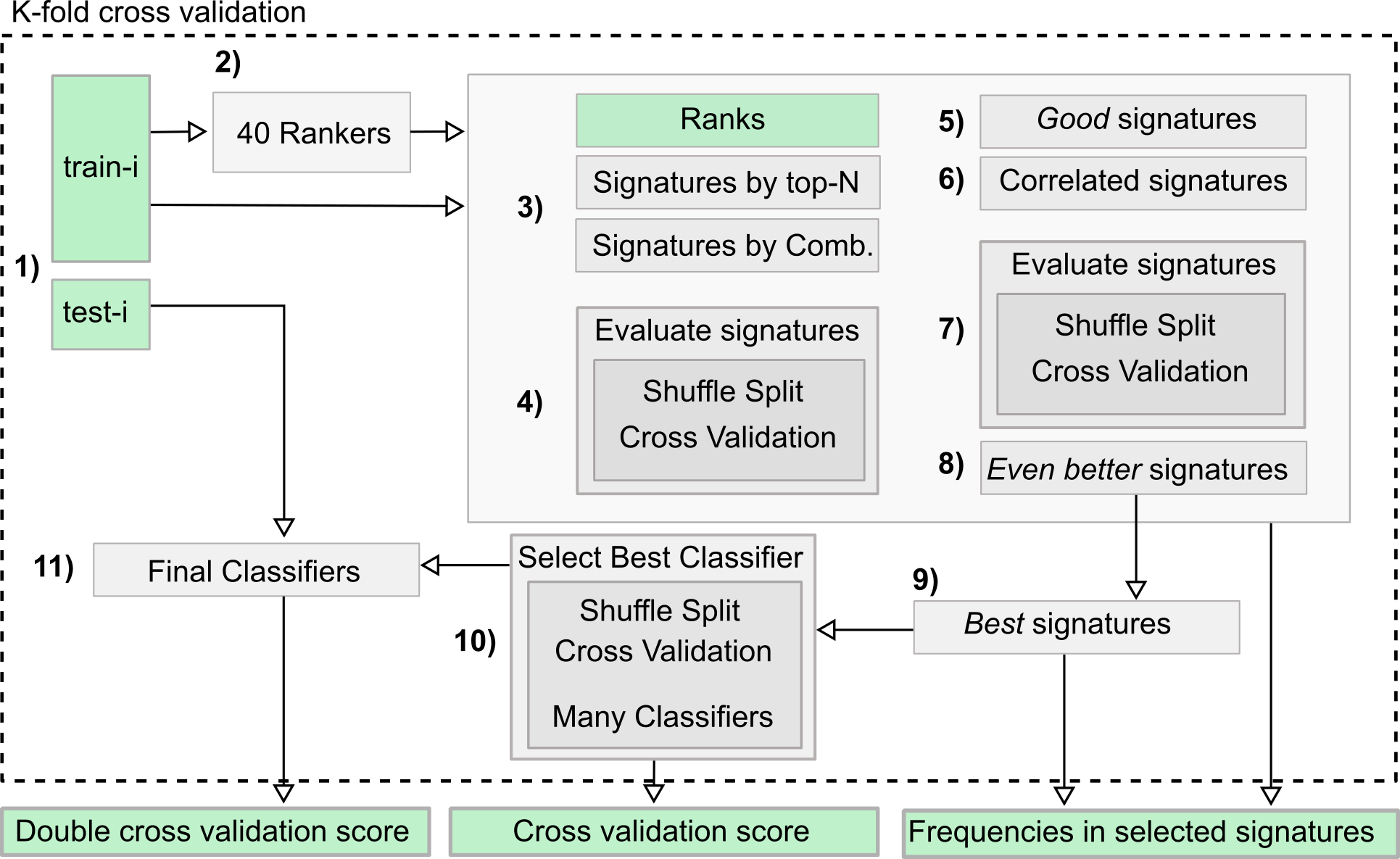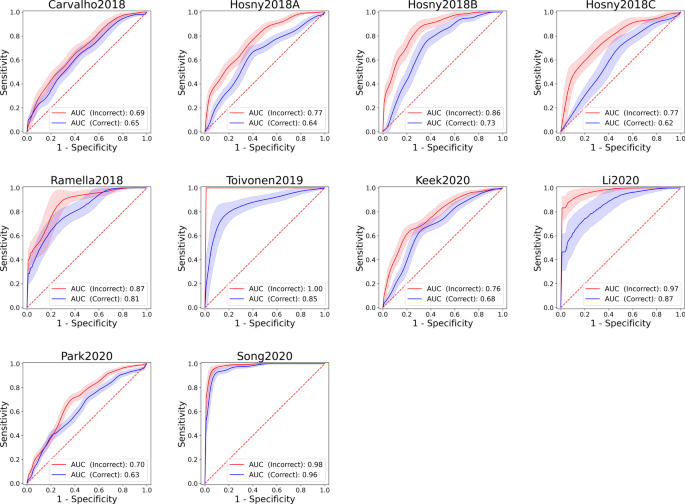
Measuring the bias of incorrect application of feature selection when using cross-validation in radiomics | Insights into Imaging | Full Text
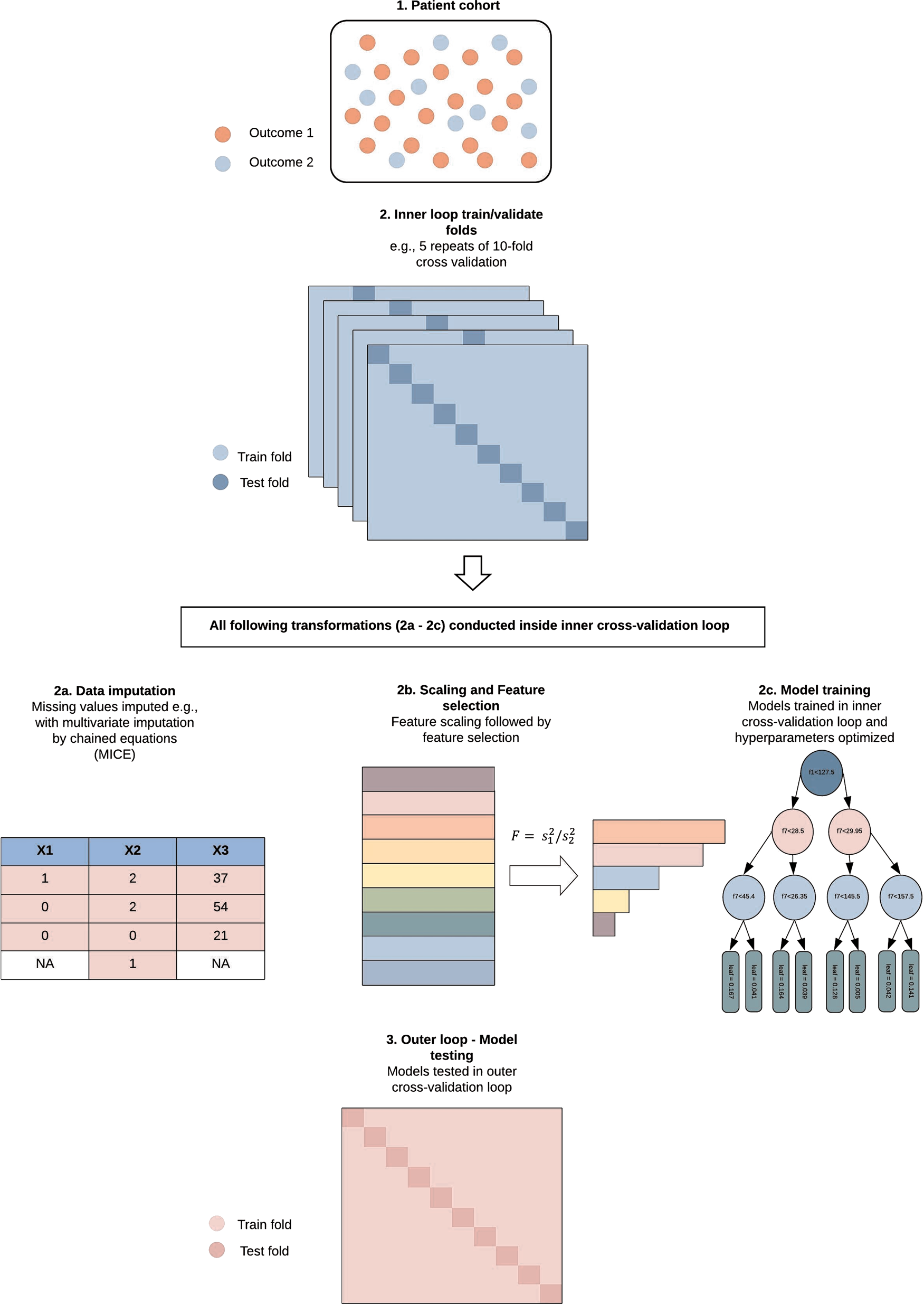
Recommendations and future directions for supervised machine learning in psychiatry | Translational Psychiatry

Hybrid Feature Selection Method Based on Neural Networks and Cross- Validation for Liver Cancer With Microarray | Semantic Scholar
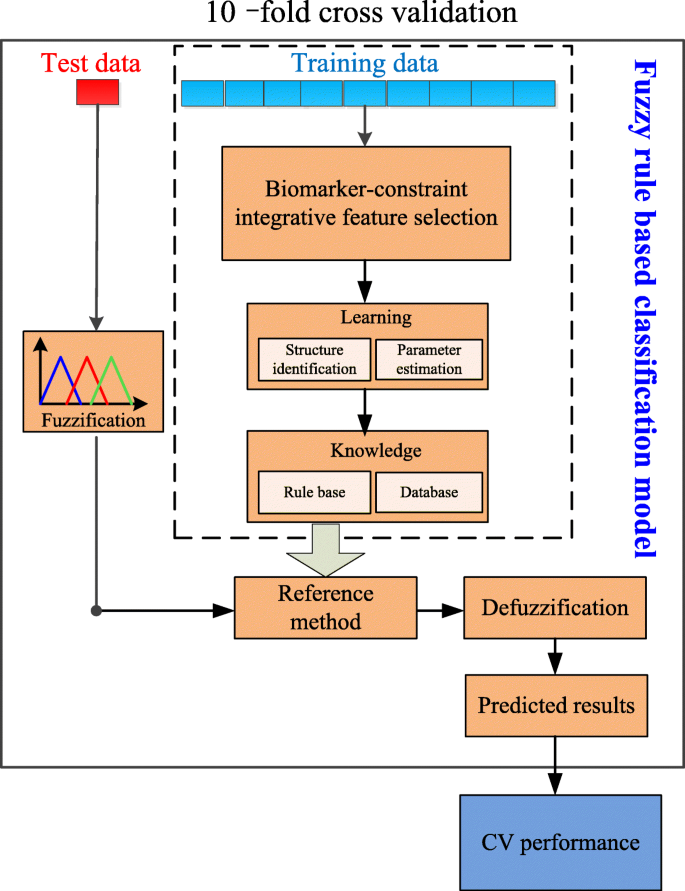
A robust fuzzy rule based integrative feature selection strategy for gene expression data in TCGA | BMC Medical Genomics | Full Text
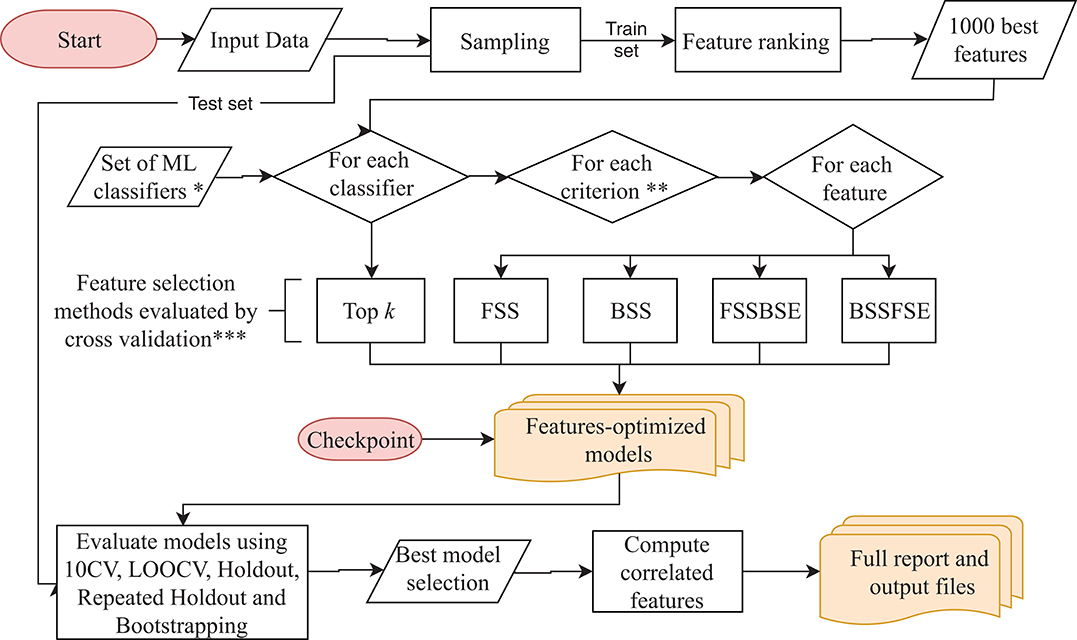
Frontiers | Large-Scale Automatic Feature Selection for Biomarker Discovery in High-Dimensional OMICs Data
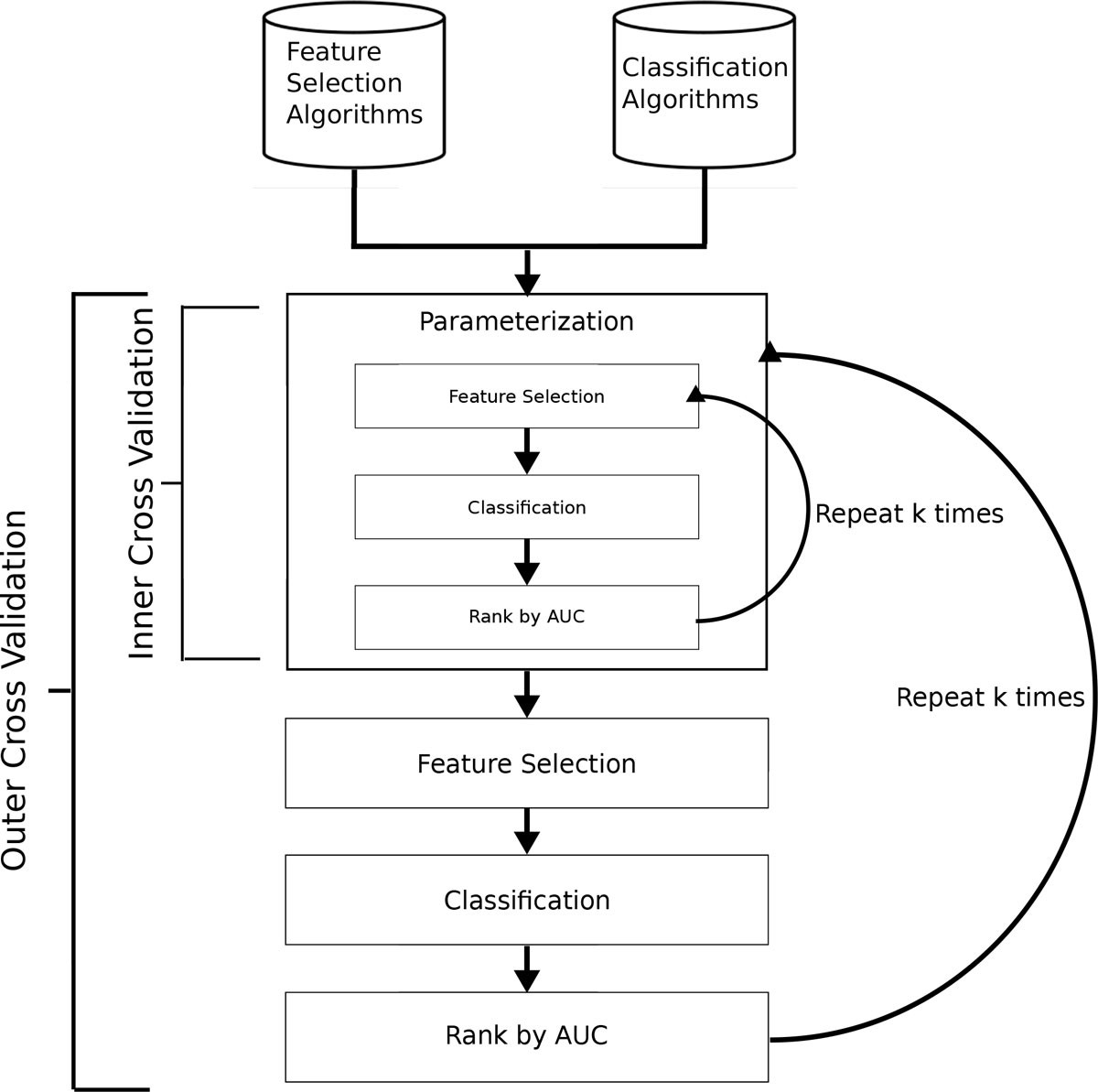
Feature selection and classifier performance on diverse bio- logical datasets | BMC Bioinformatics | Full Text
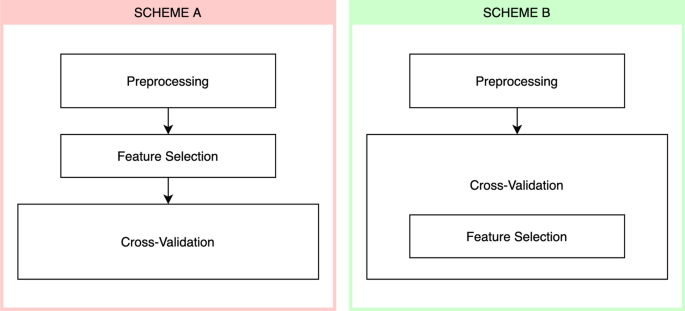
Measuring the bias of incorrect application of feature selection when using cross-validation in radiomics | Insights into Imaging | Full Text